Mankind is always on the search for who, how and why we are human. Rather uniquely as humans, we always strive for the unreachable and try to understand the impossible. Once upon a time deoxyribonucleic acid (DNA) was exactly that. As one of the most common and important structures in existence, DNA was masked in obscurity, therefore launching a fight among talented scientists to find the answers. Answers were indeed found, and in the past fifty years the level of knowledge has grown beyond expectation. In this paper I wish to deal with recently developed technologies and improvements in DNA manipulation and what this means for archaeologists. I specifically will focus on Illumina, 'a global company that develops innovative array-based solutions for DNA, RNA, and protein analysis' (Illumina 2010), as an example of the machinery and methods being employed, as well as an illustration of how commercial DNA manipulation has become.
Illumina
Upon first setting eyes on the Illumina website, I felt like I was buying a new laptop or iPod. The design and layout of the website suggests anything but analytical services and thousands of dollars worth of DNA-manipulating machines and software. The information is indeed advertising like any other technology company, and upon clicking on the systems tab, a range of shiny-looking products appear, seeming more like multimedia stations than the next step in genetic engineering. Yet that is exactly what Illumina is, a leading company in genetic engineering solutions, offering sequencing, genotyping and CNV analysis, gene regulation and epigenetic analysis, as well as PCR and genome analysing systems (Illumina 2010). Founded in 1998 by a team of highly qualified scientists and with the aid of CW Group, a venture capital firm, it now offers a total of eight systems and multiple services (Illumina 2010). Its vision is quoted as 'to be the leading provider of integrated solutions that advance the understanding of genetics and health' and its purpose 'to improve human health by enabling our customers to accelerate the collection, analysis and application of biological information' (Illumina 2010). These aims and objectives are admirable, and luckily for the archaeological world they are not limited to the healthcare service or present day biotechnology. A lack of cost efficiency may prevent utilisation of this type of software and systems for now, but if and when money is no longer an issue, archaeologists can apply this information to the past and onto ancient DNA as easily as scientists can find out information on human health and disease today via access to the human genome. This is a slow process and the systems are still far from fully accessible, yet it has potential. In this way, DNA becoming as commercial as physically possible is actually an advantage for us; the easier the techniques become and the cheaper they can be utilised, the more possibility there is for us to use such techniques in more and more archaeological questions.
PCR
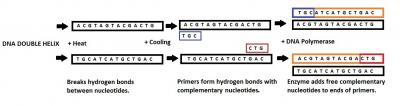
Created in 1983, PCR, or polymerase chain reaction, is a commonly employed technique (McPherson and Moller 2006). Although still extremely useful, traditional PCR has received less and less focus as other techniques have overshadowed its uses. This being said it still remains one of the most important gene manipulation techniques of our time. Figure 1 shows my own annotated picture of the PCR sequence, specifically for those unfamiliar with the idea. From its initial discovery in the 1980s, PCR has improved and changed rapidly, along the way dealing with contamination issues and technological problems. Today, as demonstrated by Illumina, standard PCR is no longer as valuable and worthwhile an application as it once was, instead the machine offered is The Eco, a real-time PCR system. Claiming to make real-time PCR available to all researchers and offering extreme sensitivity, Illumina (2010) identify that its main difference from standard PCR is that samples are measured as they are amplified rather than afterwards. What this really means is more accuracy in the whole PCR process, something that any genetic engineering company or genetic researchers have been and will continue to be searching for. This automatically launches the system into the commercial world, opens it up to a range of possible benefactors and most importantly makes it available to other disciplines such as archaeology. Although archaeology may be happy to settle for standard PCR techniques to amplify short DNA sequences, a system to make it easier, more accurate and thus potentially more cost efficient cannot be taken for granted. PCR is probably the most important development in archaeological genetics as it has allowed previously unavailable DNA to be studied. This has been most revealing in terms of ancient hominids, such as Neanderthals, where little archaeological remains are available. The process of PCR is also destructive and therefore the ability to succeed in creating the most accurate and sensitive amplification is important to ensure valuable archaeological evidence is not destroyed in vain.
DNA sequencing
The name Genome Analyzer speaks for itself. Through the use of such a machine, a genome (the complete volume of an organism's hereditary information) can be sequenced. This more often than not involves some sort of PCR to amplify the library present, again emphasising PCR's revolutionary presence in biotechnology, genetic engineering and archaeology. The use of analysing complete genomes in archaeology can be fully shown with the recent Neanderthal genome that has been draft sequenced to 40 billion nucleotides from three individuals (Affourtit et al 2010). If the availability of this service could increase, then it may be possible to progress this further, enabling us to really trace the heritage of Neanderthals and consequently modern humans more accurately than has ever been done before. It is important to note that all these techniques are surrounded by limitations, especially in terms of contamination, and although the progressions can make the DNA manipulation simpler and in principle more accurate, this will not render them error free. The Genome Analyzer IIe and Genome Analyzer IIx have a simple workflow following three steps; all based on ready to use and automated kits (Illumina 2010). The main point of these kits is to reduce hands-on time needed, an important characteristic as it reduces contamination and labour costs as well as freeing up time to focus on other things. For example, any of the sequencing systems only take approximately ten minutes to set up.
Through this paper I have given a comprehensive and insightful look into the new techniques and equipment in DNA. The extremely scientific details of the methods and techniques used in DNA manipulation have been omitted, yet these are the true wonders behind the progression in genetic engineering that has been possible. It is these principles that we should aim to utilise in archaeology where possible, and hopefully with time they will become more accessible to the archaeological field. Illumina is already a hugely successful and world renowned company pushing for developments in genetic engineering, something it is certainly achieving. The fiercely commercial world that Illumina operates in is clear whilst navigating their website pages, there is even a shopping cart for the products we may choose. Yet unlike purchasing our iPods, stereos or even cars, the scientific jargon that fills some pages highlights what commercialism sometimes detracts from, although genetic engineering may be getting cheaper and cheaper by the year it is still a hugely scientific, expensive and very customer-specific field; a field, whose incorporation into archaeological science I can only hope will continue to grow as mankind tries to find out all those who, hows and whys.
Bibliography
- Affourtit, J. Alkan, C. Aximu-Petri, A.Birney , E. Brajkovic, D. Briggs, A. Burbano, M. Butthof, A. de la Rasilla, M. Doronichev, B. Durand, E. Egholm, E. Eichler, E. Falush, D. Fortea, J. Fritz, M. Golovanova, L. Good, J. Green, R. Gušic, I. Hansen, N. Höber, B. Höffner, B. Jensen, J. Johnson , P. Kelso, J. Kircher M. Krause, J. Kucan, Z. Lachmann, M. Lalueza-Fox, C. Lander, E. Li, H. Malaspinas, A. Maricic, T. Marques-Bonet, T. Meyer, M. Mullikin, J. Nielsen, R. Novod, N. Nusbaum, C. Pääbo, S. Patterson, N. Prüfer, K. Reich, D. Rosas, A. Rudan, P. Russ, C. Schmitz, R. Schultz, R. Siegemund, M. Slatkin, M. Stenzel, U. Verna,C. Weihmann, A. Zhai, W. (2010) 'A Draft Sequence of the Neanderthal Genome' Science 328: 710-722.
- Illumina Inc. (2010) 'illumina' website. http://www.illumina.com. Page Consulted: 17/10/10.
- McPherson, M. J. and Møller, S. G. (2006) PCR. New York: Taylor and Francis Group.