Introduction
Advances in analytical methods such as the polymerase chain reaction (PCR) and next-generation sequencing (NGS) have resulted in the increasing quality and quantity of ancient DNA data in the past decade. The studies of ancient DNA have provided insights into human origins (Lazaridis et al., 2014), domestication (Librado et al., 2017), palaeopathology (Weyrich et al., 2017), and mass migration (Olalde et al., 2018). Kristiansen (2014) has highlighted that the ‘Third Scientific Revolution’ is underway, as methodological development has allowed for the direct identification of ancient genomes in a cost-effective manner.
The introduction of NGS is a major breakthrough for two reasons. Firstly, DNA fragments shorter than 90 base pairs could be sequenced (Hofman & Warinner, 2019). Secondly, NGS is less laborious than the previous combination of PCR and Sanger sequencing, as there is no need to target specific genes (van Dijk et al., 2018). After next-generation sequencers entered the market during 2007, the cost of sequencing a haploid human genome has decreased from $10 million to approximately $1000 in 2017 (Wetterstrand, 2018). If such plummet continues in the following decade, the cost of whole-genome sequencing might be reduced to almost zero. This could revolutionise the field of aDNA studies, as high-priced sequencing has been one of the primary barriers (Fernandez-Dominguez & Reynolds, 2017).
Here my focus shall be on the studies of Neolithisation throughout Eurasia in the hypothetical scenario of free whole-genome sequencing. Such focus is archaeologically significant for two reasons. Firstly, the transition from foraging to farming is one of the major dietary adaptations in human history. Transforming: human-environment relationships, demography, health, and socio-political dynamics (Fowler et al., 2015). Secondly, aDNA is one of the most direct evidence that could be used to interpret past migration and group interactions. I shall argue that the possibility of free whole-genome sequencing would impact the research of Neolithisation at three levels. At the supra-regional level, aDNA databases could be built across Eurasia. At the site level, kinship analyses could be routinely practised. At the individual level, it would be possible to reconstruct the biological characteristics of Neolithic individuals. Three caveats of aDNA analyses—the limitations of grand narratives, preservation, and contamination—will also be addressed.
Archaeological background: Neolithisation throughout Eurasia
At the outset, it is necessary to outline the process of Neolithisation across Eurasia. The general picture is that Neolithisation started around 10000 BC in the Near East (Zedar, 2011). Before c.7000 BC, Europe was a continent of foragers, while after c. 2500 BC, agriculture became the primary mode of subsistence from Greece to Ireland (Fowler et al., 2015). Such transition was large-scale and almost one-directional, as until today, most populations are still dependent on farming (Robb, 2013). Following Thomas (2013) and Bickle & Whittle (2013), the Neolithic is defined as a way of life, involving the exploitation of domesticates, sedentism, burial customs, and the use of novel material cultures. For example, pottery, stone axes, and figurines. However, such Neolithic package was seldom adopted in its entirety, nor at the same rate between c. 10000 BC and c. 2500 BC (Forenbaher & Miracle, 2005). It is clear that the emergence and duration of Neolithic cultures vary greatly from region to region throughout Eurasia.
The mechanisms involved in Neolithisation have been debated. There are two rather dichotomised models: the cultural diffusion model (CDM) and the demic diffusion model (DDM) (Ion, 2017). It is proposed in the CDM model that agriculture is spread via the exchanges of ideas and technology without the actual movement of people from the Near East into Europe (Dennell, 1983; Whittle, 1996). This implies that genetic mixing between Near Eastern farmers and indigenous European foragers is negligible. In contrast, it is postulated in the DDM model that farming is introduced by the waves of migration (Ammerman & Cavalli-Sforza, 1984). Such mass migration might lead to the large-scale replacement of the European gene pool. Between the two extremes, there are integrationist models in which certain interactions between foragers and farmers are emphasised; various mechanisms could be involved such as elite dominance (Renfrew, 1987), frontier mobility (Zvelebil & Lillie, 2000), or leapfrog colonisation (Forenbaher & Miracle, 2005).
Previously, the analyses of modern DNA have shed some light on this migration-acculturation debate. Using principal component analysis, Ammerman & Cavalli-Sforza (1984) showed that the spatial distribution of the major histocompatibility complex and mitochondrial DNA had a west-east cline in modern European populations. Such distribution was indicative of genetic trajectories due to the movement of Near Eastern farmers. However, it should be noted that mtDNA fails to represent the whole complexity of human genomes, as it is exclusively inherited from the maternal side. The use of modern gene pool could be problematic due to mutations and the loss of Neolithic signals caused by population replacement (Fernandez-Dominguez & Reynolds, 2017).
Genome-wide ancient DNA information has become available after the advancement of PCR and NGS. The analyses of ancient genomes are often in agreement with the DDM model, as the genetic make-up of European foragers appears to be homogeneous before the Neolithic transition, consisting of U-derived mitochondrial haplogroups and the I haplogroup for the Y chromosome (Haak et al., 2015; Lazaridis et al., 2014; Mathieson et al., 2015; Mathieson et al., 2018). However, there are two major issues: small sample size and limited data variability. So far, fewer than 500 ancient genomes have been sequenced, with central European data predominant (Lipson et al., 2017). Overall, DNA analyses have established that mass migration might occur during the Neolithic, while its timing, extent, and transmission at different scales have not been fully understood.
Supra-regional level: the development of aDNA databases
Back to the hypothetical scenario of free genome-wide sequencing, the first impact is likely to be the development of large-scale, spatial-temporal aDNA databases. As previously mentioned, while certain regions such as Saxony-Anhalt in Germany have been intensively investigated, other key areas, for example, the Near East, the Balkans, and Greece, remain poorly studied (Lipson et al., 2017). There are at least five major domestication centres in the Near East: Anatolia, south-eastern Turkey, Iraq, northern Syria, and the Levant (Zedar, 2011). However, genome-wide ancient DNA data has only been obtained from only 41 individuals in these areas, including the Levant, the Zagros mountains in Western Iran, and central Anatolia (Broushaki et al., 2016; Lazaridis et al., 2016). Hence, a wider and more representative sampling is required to capture the whole picture of past populations and the routes of expansion from the Near East into Europe.
The establishment of aDNA databases might contribute to the understanding of Neolithisation in three aspects. Firstly, in terms of the Near East, large-scale, genome-wide aDNA data would elucidate the genetic make-up of Near Eastern farmers from Pre-Pottery Neolithic A (c.8500- c.7600 BC) to Pre Pottery Neolithic C (c.6000- c.5500 BC) (Zedar, 2011). This is essential in determining noise-to-signal ratios in order to distinguish the input of Near Eastern genes from the European genetic background. Secondly, regarding frontier zones such as the Aegean and the Balkans, the establishment of aDNA databases would elucidate the magnitude of migration and the degree of interactions. Thirdly, it would be possible to narrow down the time frame of migratory events throughout Europe using high-resolution aDNA databases.
Site level: the routine assessments of kinships
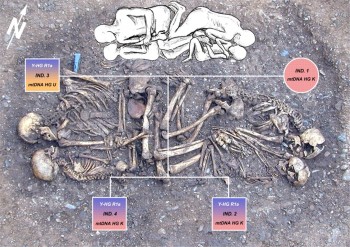
Moving on to the site level, free whole-genome sequencing could allow for the routine assessments of kinships. It is often challenging to determine the genetic affinity of individuals at a cemetery based on the arrangements of grave goods, burial positions, or non-metric skeletal traits (Brown & Brown, 2011). aDNA could provide direct evidence for kinships, as the chromosomes of children are inherited from their parents. One of the most emphatic examples is a parent-child burial at Eulau, Germany, dating to 2700-2400 cal BC (Haak et al., 2008). One female (35-50 years old), one male (40-60 years old), and two juveniles were buried in the same grave (see Figure 1). The adults were buried in a flexed position, facing their children. The results of aDNA analyses show that the woman and the two juveniles share the same haplogroup K1b, and all the three males have the same Y chromosome haplogroup R1a. Hence, it is evident that the four individuals buried at Eulau are genetically related to each other.
aDNA analyses have seldom been used to identify genetic relationships at early Neolithic cemeteries. The previous Eulau example dates to the Corded Ware Culture (c. 2900- c. 2400 cal BC) (Haak et al., 2008). It should also be noted that the aDNA of the Eulau individuals was analysed using PCR, while the length of most Neolithic DNA might be shorter than 90 base pairs (Kuhn et al., 2018). If the cost of whole-genome sequencing reaches zero, NGS could be applied routinely on Neolithic individuals. This is advantageous in studying Neolithisation, as genetic relationships are important in understanding personal identities, group interactions, and martial patterns such as exogamy (Brown & Brown, 2011).
Individual level: the routine reconstruction of biological characteristics
Lastly, as the cost of aDNA sequencing continues to approach zero, it would be possible to routinely reconstruct the biological characteristics of an individual, for example, eye colour, skin pigmentation, height potential, and lactase persistence. Thus far, such reconstruction has been limited to exceptionally well-preserved human remains such as the Ice Man (Keller et al., 2012). NGS has allowed for the sequencing of short and less complex aDNA, which is supported by the phenotypic reconstruction of an early Neolithic female from the Zagros Mountains of Iran (Gallego-Llorente et al., 2016). This female was buried under a mound dating to 9650-9950 cal BP, and her age at death is estimated to be between 30 and 50 years old. Her petrous bone was extracted and sequenced. Based on diagnostics single nucleotide polymorphisms (SNPs), she is most likely to have dark skin, black hair, and brown eyes. Although the derived variant of the SLC45A2 gene, which is associated with light skin pigmentation, is absent, she had at least one copy of the derived SLC24A5 allele which is common amongst Anatolian farmers. The absence of the LCT gene suggests that she was lactose-intolerant. Overall, it is possible to obtain detailed biological information using aDNA at the individual level. If more phenotypic data become available due to the reduced cost of aDNA sequencing, this might affect the studies of Neolithisation in three ways. Firstly, the temporal-spatial distribution of biological characteristics throughout Neolithic Eurasia could be elucidated. Secondly, it would allow for the facial reconstruction of Near Eastern farmers and European foragers. Appearance is integral to identities, which might affect interactions between populations if there are visual differences (Jones & Graves-Brown, 2013). Lastly, phenotypes might be affected by the admixture of populations. It would be possible to investigate their transmission and introgression during the process of Neolithisation with increased data.
Caveats & Mitigation
It is evident from the previous discussion that in the hypothesised scenario of free aDNA sequencing, the studies of Neolithisation throughout Eurasia might be impacted at three levels: supra-regional, site and individual. However, three caveats should be considered: the limitations of grand narratives, preservation, and contamination. The first caveat is of epistemological significance. The interpretations of aDNA analyses are often criticised as grand narratives which undermine the contributions of internal social dynamics such as agency (Heyd, 2017). Indeed grand narratives seemed to be disregarded during the heyday of post-processualism, given the diversity and context-specificity of social changes (Hodder, 2003). However, it should be acknowledged that one key strength of archaeological research is its spatial-temporal depth. Back to the debate of the Neolithisation process, it is arguably most evident on a supra-regional level in the longue durée, as most European populations practised farming after c.2500 BC (Robb, 2013).
Secondly, aDNA preservation is largely restricted by time and burial environments such as temperature, pH, and hydrology (Brown & Brown, 2011). Although the cost of aDNA sequencing could be reduced to almost zero, the availability of aDNA data is still limited. For example, aDNA data from the Near East or the Mediterranean is scarce due to their high average temperatures (Fernandez-Dominguez & Reynolds, 2017). However, this issue could be mitigated by hybridisation capture-based target enrichment (Olalde et al., 2018) or using substrates that tend to yield high endogenous contents such as petrous bones (Gallego-Llorente et al., 2016). Lastly, aDNA could be contaminated by various processes during excavation and post-excavation analyses. This could be mitigated by following the established protocols to minimise contamination in laboratories and in the field (Hofman & Warinner, 2019).
Concluding remarks
If the cost of aDNA sequencing becomes free in the next decade, the studies of Neolithisation might be affected at three scales. At the supra-regional level, large-scale, high-resolution databases might be developed to elucidate the spatial-temporal variations of past genomes throughout Eurasia. At the site level, analysing kinship using aDNA data could be a routine practice, contributing to the understanding of social organisation, martial patterns, and group interactions. At the individual level, the biological characteristics of Neolithic individuals could be routinely reconstructed by aDNA studies. This might also shed some light on the origins and transmission of phenotypes. Although the interpretations of aDNA analyses could be restricted by issues such as preservation and contamination, these could be mitigated by technological development and the increased awareness of epistemological and methodological limitations.






